Imaging Software Rapidly Identifies Dendritic Spines
|
By MedImaging International staff writers Posted on 17 Jul 2018 |
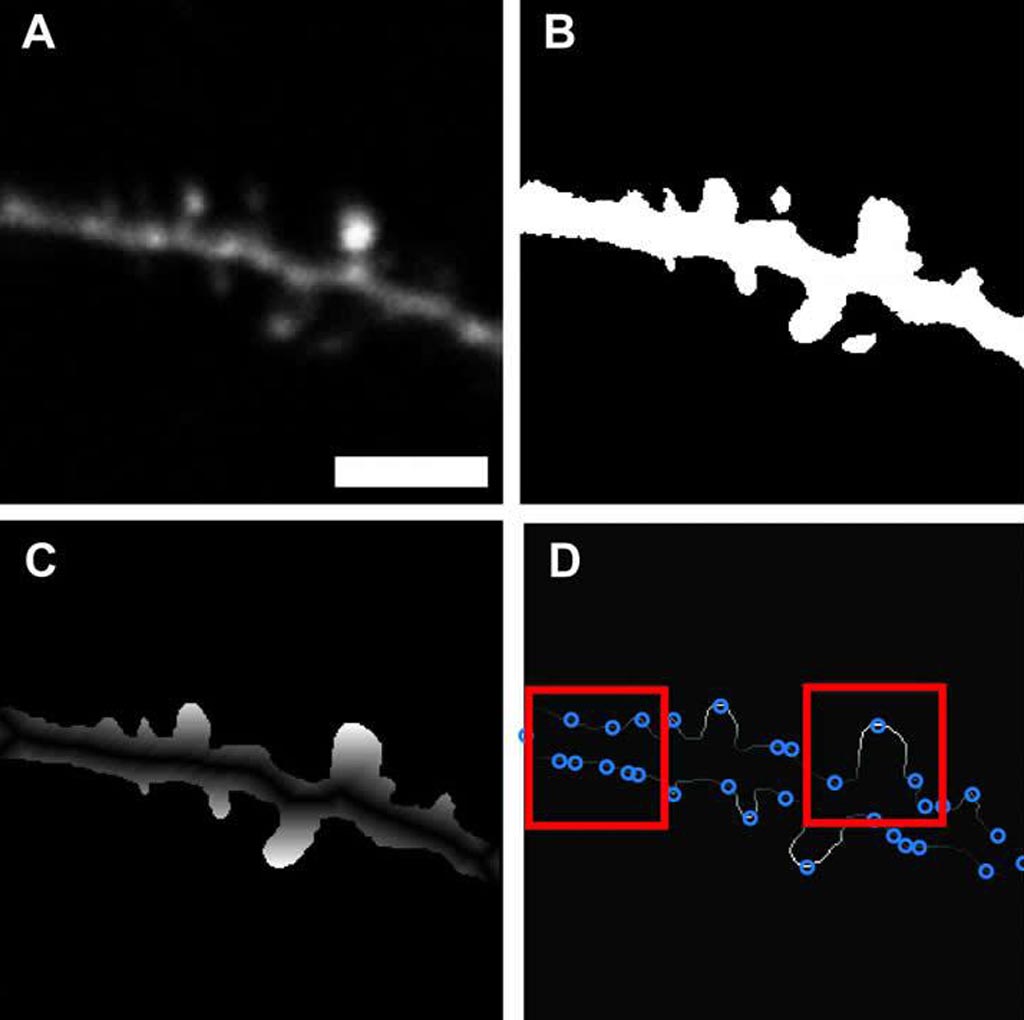
Image: Surface smoothing of a binary image of the dendritic spine (Photo courtesy of Max Planck Florida Institute).
A novel 2-photon microscopy algorithm autonomously and efficiently identifies small dendritic spines with over 90% accuracy, according to a new study.
Developed by researchers at the Max Planck Florida Institute for Neuroscience (Jupiter, FL; USA), the software is built on a machine learning approach that uses custom thresholding and binarization functions to clean up fluorescent images, and a neural network trained to differentiate between the dendrite backbone and dendritic spines using the relative shape of the spine perimeter and its corresponding dendritic backbone. Once the training period finishes, the software can automatically scan through an image and demarcate spines it comes across with high precision.
By automating the process of spine identification, the high-throughput imaging system has the potential to dramatically increase workflow efficiency by stimulating and monitoring hundreds of dendritic spines under various conditions at the same time, shaving hours off analysis time. In addition, the source code is provided free of charge bundled together with a user-friendly, open-source, MATLAB-based software package for spine analysis. The study was published on July 5, 2018, in PLOS One.
“When engineers and scientists design cutting-edge microscopes, they usually focus on the actual physical components and design. They are mostly interested in what these imaging technologies can do, what boundaries they can break and how they perform,” said lead author Michael Smirnov, PhD. “Far less attention is placed on how these complex technologies can be made accessible for the average user and really improve their workflow. Each time I write my software, I always think about the user first; how can I make a difference for the person using it, make their research a little easier.”
Synaptic plasticity, the cellular basis for learning and memory, is mediated by a complex biochemical network of signaling proteins. These proteins are compartmentalized in dendritic spines, the tiny, bulbous, post-synaptic structures found on neuronal dendrites.
Related Links:
Max Planck Florida Institute for Neuroscience
Developed by researchers at the Max Planck Florida Institute for Neuroscience (Jupiter, FL; USA), the software is built on a machine learning approach that uses custom thresholding and binarization functions to clean up fluorescent images, and a neural network trained to differentiate between the dendrite backbone and dendritic spines using the relative shape of the spine perimeter and its corresponding dendritic backbone. Once the training period finishes, the software can automatically scan through an image and demarcate spines it comes across with high precision.
By automating the process of spine identification, the high-throughput imaging system has the potential to dramatically increase workflow efficiency by stimulating and monitoring hundreds of dendritic spines under various conditions at the same time, shaving hours off analysis time. In addition, the source code is provided free of charge bundled together with a user-friendly, open-source, MATLAB-based software package for spine analysis. The study was published on July 5, 2018, in PLOS One.
“When engineers and scientists design cutting-edge microscopes, they usually focus on the actual physical components and design. They are mostly interested in what these imaging technologies can do, what boundaries they can break and how they perform,” said lead author Michael Smirnov, PhD. “Far less attention is placed on how these complex technologies can be made accessible for the average user and really improve their workflow. Each time I write my software, I always think about the user first; how can I make a difference for the person using it, make their research a little easier.”
Synaptic plasticity, the cellular basis for learning and memory, is mediated by a complex biochemical network of signaling proteins. These proteins are compartmentalized in dendritic spines, the tiny, bulbous, post-synaptic structures found on neuronal dendrites.
Related Links:
Max Planck Florida Institute for Neuroscience
Latest General/Advanced Imaging News
- PET Scans Reveal Hidden Inflammation in Multiple Sclerosis Patients
- Artificial Intelligence Evaluates Cardiovascular Risk from CT Scans
- New AI Method Captures Uncertainty in Medical Images
- CT Coronary Angiography Reduces Need for Invasive Tests to Diagnose Coronary Artery Disease
- Novel Blood Test Could Reduce Need for PET Imaging of Patients with Alzheimer’s
- CT-Based Deep Learning Algorithm Accurately Differentiates Benign From Malignant Vertebral Fractures
- Minimally Invasive Procedure Could Help Patients Avoid Thyroid Surgery
- Self-Driving Mobile C-Arm Reduces Imaging Time during Surgery
- AR Application Turns Medical Scans Into Holograms for Assistance in Surgical Planning
- Imaging Technology Provides Ground-Breaking New Approach for Diagnosing and Treating Bowel Cancer
- CT Coronary Calcium Scoring Predicts Heart Attacks and Strokes
- AI Model Detects 90% of Lymphatic Cancer Cases from PET and CT Images
- Breakthrough Technology Revolutionizes Breast Imaging
- State-Of-The-Art System Enhances Accuracy of Image-Guided Diagnostic and Interventional Procedures
- Catheter-Based Device with New Cardiovascular Imaging Approach Offers Unprecedented View of Dangerous Plaques
- AI Model Draws Maps to Accurately Identify Tumors and Diseases in Medical Images
Channels
Radiography
view channel
Novel Breast Imaging System Proves As Effective As Mammography
Breast cancer remains the most frequently diagnosed cancer among women. It is projected that one in eight women will be diagnosed with breast cancer during her lifetime, and one in 42 women who turn 50... Read more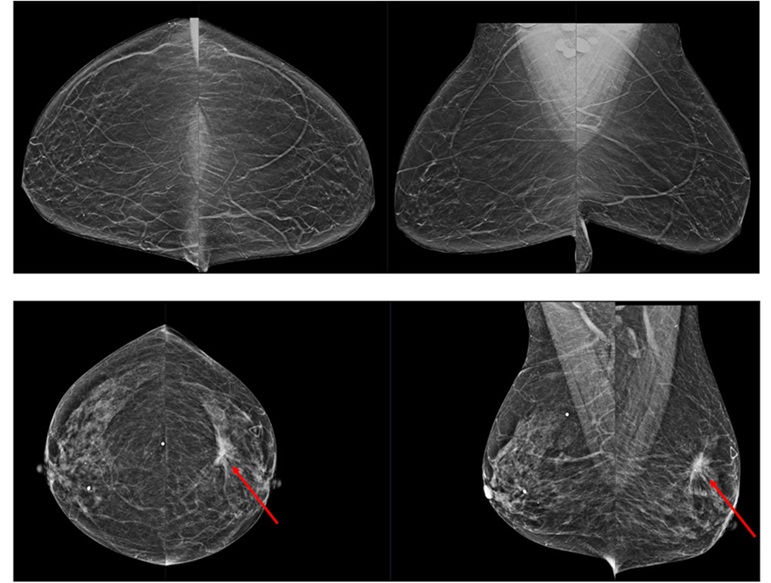
AI Assistance Improves Breast-Cancer Screening by Reducing False Positives
Radiologists typically detect one case of cancer for every 200 mammograms reviewed. However, these evaluations often result in false positives, leading to unnecessary patient recalls for additional testing,... Read moreMRI
view channel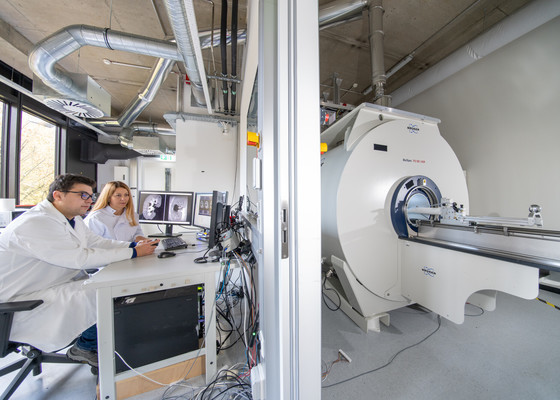
Diamond Dust Could Offer New Contrast Agent Option for Future MRI Scans
Gadolinium, a heavy metal used for over three decades as a contrast agent in medical imaging, enhances the clarity of MRI scans by highlighting affected areas. Despite its utility, gadolinium not only... Read more.jpg)
Combining MRI with PSA Testing Improves Clinical Outcomes for Prostate Cancer Patients
Prostate cancer is a leading health concern globally, consistently being one of the most common types of cancer among men and a major cause of cancer-related deaths. In the United States, it is the most... Read more
PET/MRI Improves Diagnostic Accuracy for Prostate Cancer Patients
The Prostate Imaging Reporting and Data System (PI-RADS) is a five-point scale to assess potential prostate cancer in MR images. PI-RADS category 3 which offers an unclear suggestion of clinically significant... Read more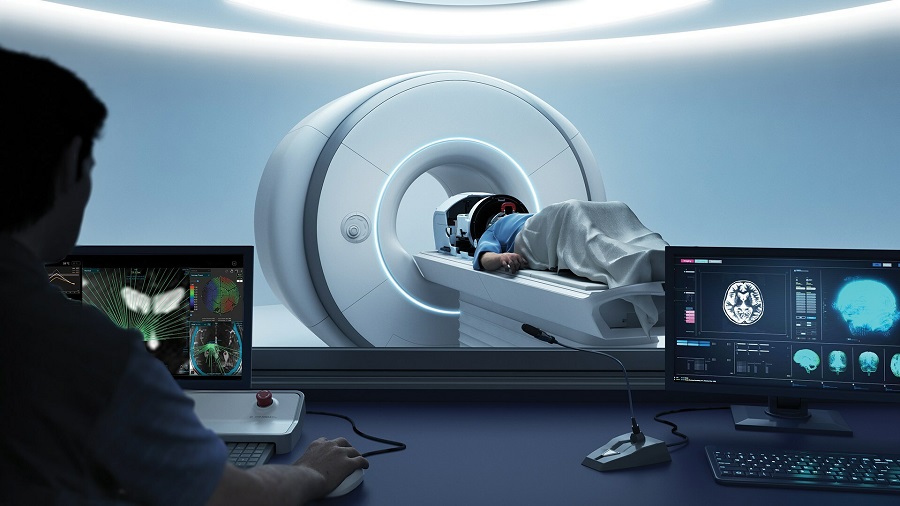
Next Generation MR-Guided Focused Ultrasound Ushers In Future of Incisionless Neurosurgery
Essential tremor, often called familial, idiopathic, or benign tremor, leads to uncontrollable shaking that significantly affects a person’s life. When traditional medications do not alleviate symptoms,... Read moreUltrasound
view channel.jpg)
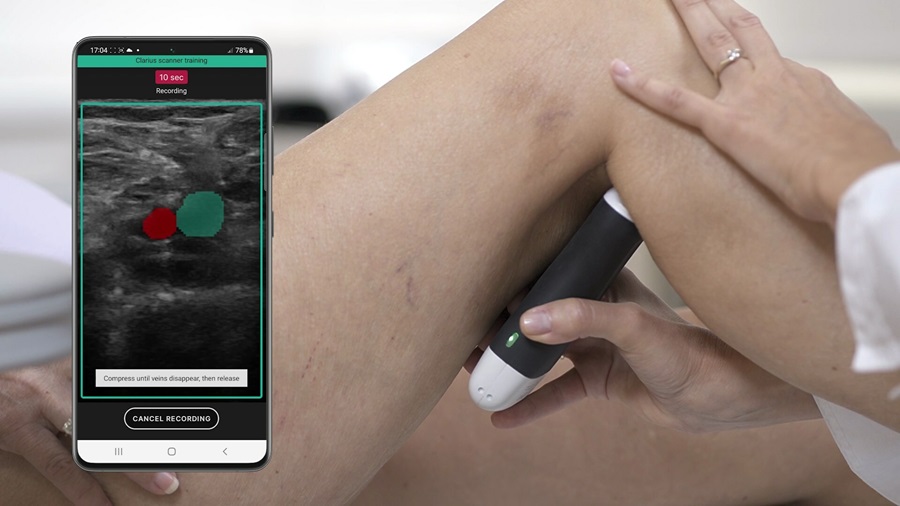
Groundbreaking Technology Enables Precise, Automatic Measurement of Peripheral Blood Vessels
The current standard of care of using angiographic information is often inadequate for accurately assessing vessel size in the estimated 20 million people in the U.S. who suffer from peripheral vascular disease.... Read more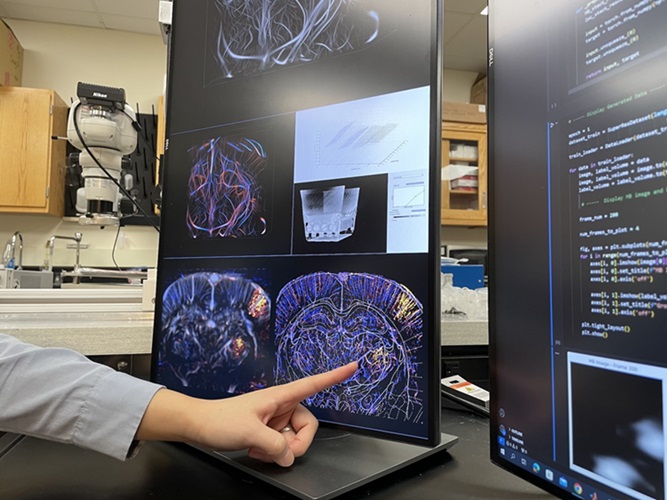
Deep Learning Advances Super-Resolution Ultrasound Imaging
Ultrasound localization microscopy (ULM) is an advanced imaging technique that offers high-resolution visualization of microvascular structures. It employs microbubbles, FDA-approved contrast agents, injected... Read more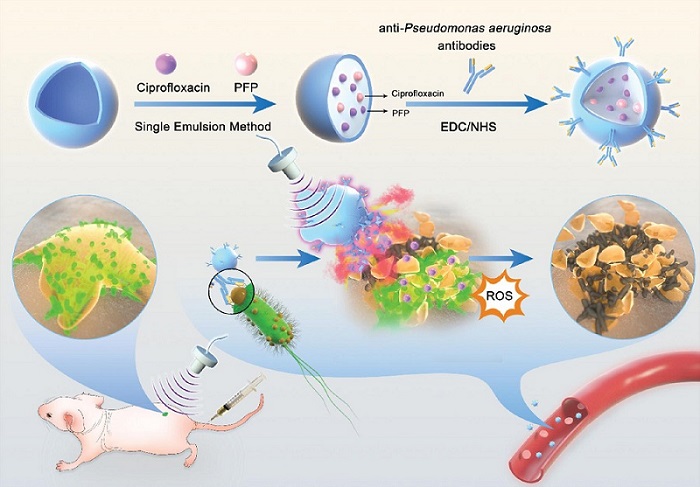
Novel Ultrasound-Launched Targeted Nanoparticle Eliminates Biofilm and Bacterial Infection
Biofilms, formed by bacteria aggregating into dense communities for protection against harsh environmental conditions, are a significant contributor to various infectious diseases. Biofilms frequently... Read moreNuclear Medicine
view channel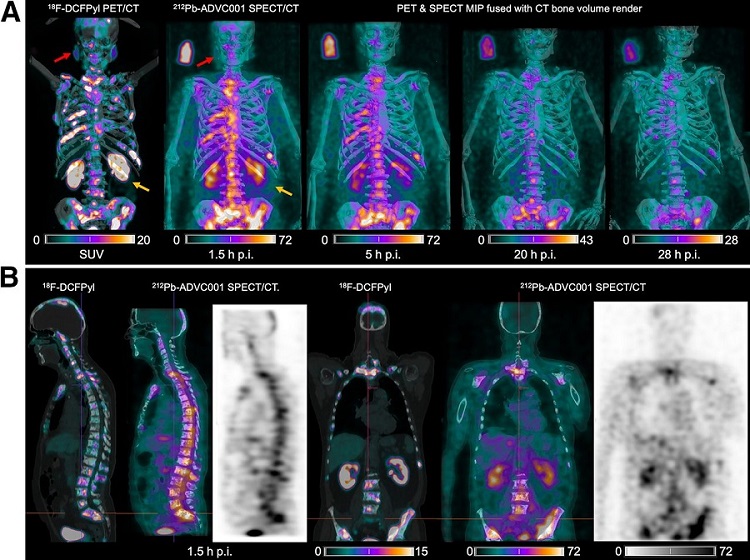
New SPECT/CT Technique Could Change Imaging Practices and Increase Patient Access
The development of lead-212 (212Pb)-PSMA–based targeted alpha therapy (TAT) is garnering significant interest in treating patients with metastatic castration-resistant prostate cancer. The imaging of 212Pb,... Read moreNew Radiotheranostic System Detects and Treats Ovarian Cancer Noninvasively
Ovarian cancer is the most lethal gynecological cancer, with less than a 30% five-year survival rate for those diagnosed in late stages. Despite surgery and platinum-based chemotherapy being the standard... Read more
AI System Automatically and Reliably Detects Cardiac Amyloidosis Using Scintigraphy Imaging
Cardiac amyloidosis, a condition characterized by the buildup of abnormal protein deposits (amyloids) in the heart muscle, severely affects heart function and can lead to heart failure or death without... Read moreImaging IT
view channel
New Google Cloud Medical Imaging Suite Makes Imaging Healthcare Data More Accessible
Medical imaging is a critical tool used to diagnose patients, and there are billions of medical images scanned globally each year. Imaging data accounts for about 90% of all healthcare data1 and, until... Read more
Global AI in Medical Diagnostics Market to Be Driven by Demand for Image Recognition in Radiology
The global artificial intelligence (AI) in medical diagnostics market is expanding with early disease detection being one of its key applications and image recognition becoming a compelling consumer proposition... Read moreIndustry News
view channel
Bayer and Google Partner on New AI Product for Radiologists
Medical imaging data comprises around 90% of all healthcare data, and it is a highly complex and rich clinical data modality and serves as a vital tool for diagnosing patients. Each year, billions of medical... Read more