Computer Program Bests Radiologists at Analyzing Brain MRI
|
By MedImaging International staff writers Posted on 27 Sep 2016 |
Image: A computer program beats radiologists in MRI analysis (Photo courtesy of CWRU).
A new study that matched two physicians against a computer algorithm in analysis of magnetic resonance imaging (MRI) brain scans found the program was nearly twice as accurate.
Researchers at Case Western Reserve University (CWRU; Cleveland, OH, USA) and the University of Texas Southwestern Medical Center (Dallas, TX, USA) conducted a study to determine the feasibility of using computer-extracted texture features to differentiate between radiation necrosis and recurrent brain tumors on post-radiochemotherapy MRI scans. In all, 58 patient scans were used, with 43 forming the training cohort for the algorithm and 15 forming the test cohort.
A set of radiomic features was extracted for every lesion on each MRI sequence - gadolinium T1WI, T2WI, and FLAIR. Feature selection was used to identify the top five most discriminating features for every MRI sequence on the training cohort. These features were then evaluated on the test cohort by a support vector machine classifier. The classification performance was compared against diagnostic reads by two expert neuroradiologists, who had access to the same MRI sequences as the classifier. Finally, clinical histologic findings were confirmed by an experienced neuropathologist.
The results revealed that in the direct comparison, one neuroradiologist diagnosed seven patients correctly, and the second physician correctly diagnosed eight patients. The computer program, on the other hand, was correct on 12 of the 15 MRI scans. The researchers are now seeking to validate the algorithms' accuracy using a much larger collection of images from across different sites, so that it could eventually be used as a decision support tool to assist neuroradiologists in improving their confidence in identifying a suspicious lesion. The study was published on September 15, 2016, in the American Journal of Neuroradiology.
“One of the biggest challenges with the evaluation of brain tumor treatment is distinguishing between the confounding effects of radiation and cancer recurrence; on an MRI, they look very similar,” said lead author biomedical engineer Pallavi Tiwari, PhD, of CWRU. “What the algorithms see that the radiologists don't are the subtle differences in quantitative measurements of tumor heterogeneity and breakdown in microarchitecture on MRI, which are higher for tumor recurrence.”
“While the physicians use the intensity of pixels on MRI scans as a guide, the computer looks at the edges of each pixel; if the edges all point to the same direction, the architecture is preserved,” added senior author professor of biomedical engineering Anant Madabhushi, PHD, director of the CWRU Center of Computational Imaging and Personalized Diagnostics. “If they point in different directions, the architecture is disrupted—the entropy, or disorder, and heterogeneity are higher.”
In a recent competition at the 2016 International Symposium of Biomedical Imaging (ISBI), held during April in Prague (Czech Republic), a machine-learning computer algorithm that was trained to recognize breast cancer metastasis in lymph nodes identified correctly 92% percent of the time, nearly matching the 96% success rate of a human pathologist.
Related Links:
Case Western Reserve University
University of Texas Southwestern Medical Center
Researchers at Case Western Reserve University (CWRU; Cleveland, OH, USA) and the University of Texas Southwestern Medical Center (Dallas, TX, USA) conducted a study to determine the feasibility of using computer-extracted texture features to differentiate between radiation necrosis and recurrent brain tumors on post-radiochemotherapy MRI scans. In all, 58 patient scans were used, with 43 forming the training cohort for the algorithm and 15 forming the test cohort.
A set of radiomic features was extracted for every lesion on each MRI sequence - gadolinium T1WI, T2WI, and FLAIR. Feature selection was used to identify the top five most discriminating features for every MRI sequence on the training cohort. These features were then evaluated on the test cohort by a support vector machine classifier. The classification performance was compared against diagnostic reads by two expert neuroradiologists, who had access to the same MRI sequences as the classifier. Finally, clinical histologic findings were confirmed by an experienced neuropathologist.
The results revealed that in the direct comparison, one neuroradiologist diagnosed seven patients correctly, and the second physician correctly diagnosed eight patients. The computer program, on the other hand, was correct on 12 of the 15 MRI scans. The researchers are now seeking to validate the algorithms' accuracy using a much larger collection of images from across different sites, so that it could eventually be used as a decision support tool to assist neuroradiologists in improving their confidence in identifying a suspicious lesion. The study was published on September 15, 2016, in the American Journal of Neuroradiology.
“One of the biggest challenges with the evaluation of brain tumor treatment is distinguishing between the confounding effects of radiation and cancer recurrence; on an MRI, they look very similar,” said lead author biomedical engineer Pallavi Tiwari, PhD, of CWRU. “What the algorithms see that the radiologists don't are the subtle differences in quantitative measurements of tumor heterogeneity and breakdown in microarchitecture on MRI, which are higher for tumor recurrence.”
“While the physicians use the intensity of pixels on MRI scans as a guide, the computer looks at the edges of each pixel; if the edges all point to the same direction, the architecture is preserved,” added senior author professor of biomedical engineering Anant Madabhushi, PHD, director of the CWRU Center of Computational Imaging and Personalized Diagnostics. “If they point in different directions, the architecture is disrupted—the entropy, or disorder, and heterogeneity are higher.”
In a recent competition at the 2016 International Symposium of Biomedical Imaging (ISBI), held during April in Prague (Czech Republic), a machine-learning computer algorithm that was trained to recognize breast cancer metastasis in lymph nodes identified correctly 92% percent of the time, nearly matching the 96% success rate of a human pathologist.
Related Links:
Case Western Reserve University
University of Texas Southwestern Medical Center
Latest MRI News
- Low-Cost Whole-Body MRI Device Combined with AI Generates High-Quality Results
- World's First Whole-Body Ultra-High Field MRI Officially Comes To Market
- World's First Sensor Detects Errors in MRI Scans Using Laser Light and Gas
- Diamond Dust Could Offer New Contrast Agent Option for Future MRI Scans
- Combining MRI with PSA Testing Improves Clinical Outcomes for Prostate Cancer Patients
- PET/MRI Improves Diagnostic Accuracy for Prostate Cancer Patients
- Next Generation MR-Guided Focused Ultrasound Ushers In Future of Incisionless Neurosurgery
- Two-Part MRI Scan Detects Prostate Cancer More Quickly without Compromising Diagnostic Quality
- World’s Most Powerful MRI Machine Images Living Brain with Unrivaled Clarity
- New Whole-Body Imaging Technology Makes It Possible to View Inflammation on MRI Scan
- Combining Prostate MRI with Blood Test Can Avoid Unnecessary Prostate Biopsies
- New Treatment Combines MRI and Ultrasound to Control Prostate Cancer without Serious Side Effects
- MRI Improves Diagnosis and Treatment of Prostate Cancer
- Combined PET-MRI Scan Improves Treatment for Early Breast Cancer Patients
- 4D MRI Could Improve Clinical Assessment of Heart Blood Flow Abnormalities
- MRI-Guided Focused Ultrasound Therapy Shows Promise in Treating Prostate Cancer
Channels
Radiography
view channel
Novel Breast Imaging System Proves As Effective As Mammography
Breast cancer remains the most frequently diagnosed cancer among women. It is projected that one in eight women will be diagnosed with breast cancer during her lifetime, and one in 42 women who turn 50... Read more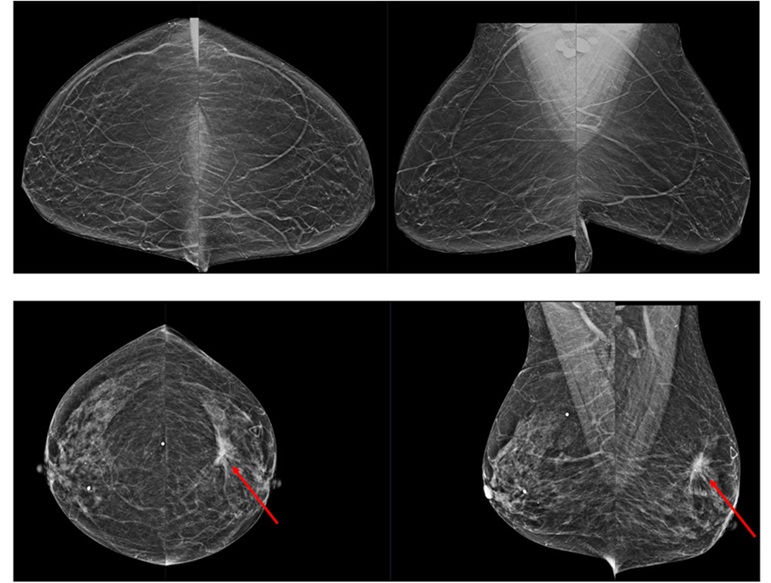
AI Assistance Improves Breast-Cancer Screening by Reducing False Positives
Radiologists typically detect one case of cancer for every 200 mammograms reviewed. However, these evaluations often result in false positives, leading to unnecessary patient recalls for additional testing,... Read moreUltrasound
view channel.jpg)
Diagnostic System Automatically Analyzes TTE Images to Identify Congenital Heart Disease
Congenital heart disease (CHD) is one of the most prevalent congenital anomalies worldwide, presenting substantial health and financial challenges for affected patients. Early detection and treatment of... Read more
Super-Resolution Imaging Technique Could Improve Evaluation of Cardiac Conditions
The heart depends on efficient blood circulation to pump blood throughout the body, delivering oxygen to tissues and removing carbon dioxide and waste. Yet, when heart vessels are damaged, it can disrupt... Read more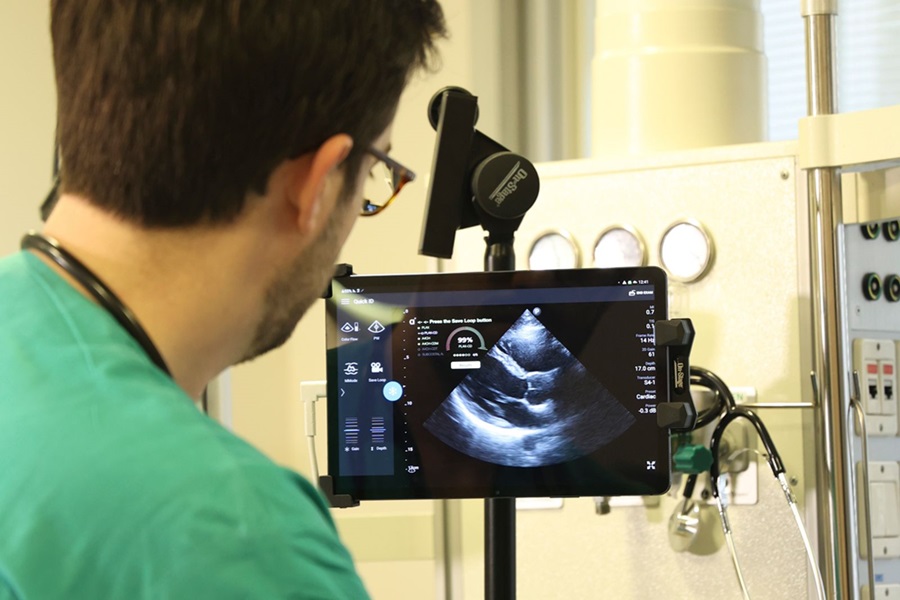
First AI-Powered POC Ultrasound Diagnostic Solution Helps Prioritize Cases Based On Severity
Ultrasound scans are essential for identifying and diagnosing various medical conditions, but often, patients must wait weeks or months for results due to a shortage of qualified medical professionals... Read moreNuclear Medicine
view channel
New PET Biomarker Predicts Success of Immune Checkpoint Blockade Therapy
Immunotherapies, such as immune checkpoint blockade (ICB), have shown promising clinical results in treating melanoma, non-small cell lung cancer, and other tumor types. However, the effectiveness of these... Read moreNew PET Agent Rapidly and Accurately Visualizes Lesions in Clear Cell Renal Cell Carcinoma Patients
Clear cell renal cell cancer (ccRCC) represents 70-80% of renal cell carcinoma cases. While localized disease can be effectively treated with surgery and ablative therapies, one-third of patients either... Read more
New Imaging Technique Monitors Inflammation Disorders without Radiation Exposure
Imaging inflammation using traditional radiological techniques presents significant challenges, including radiation exposure, poor image quality, high costs, and invasive procedures. Now, new contrast... Read more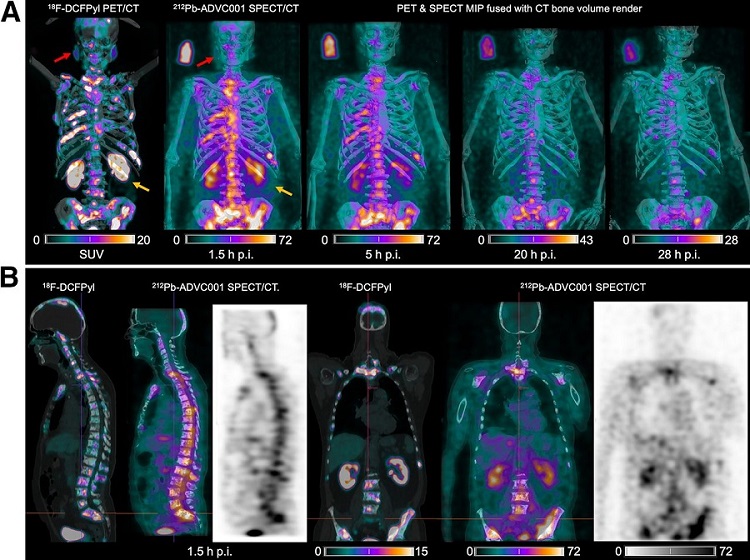
New SPECT/CT Technique Could Change Imaging Practices and Increase Patient Access
The development of lead-212 (212Pb)-PSMA–based targeted alpha therapy (TAT) is garnering significant interest in treating patients with metastatic castration-resistant prostate cancer. The imaging of 212Pb,... Read moreGeneral/Advanced Imaging
view channelBone Density Test Uses Existing CT Images to Predict Fractures
Osteoporotic fractures are not only devastating and deadly, especially hip fractures, but also impose significant costs. They rank among the top chronic diseases in terms of disability-adjusted life years... Read more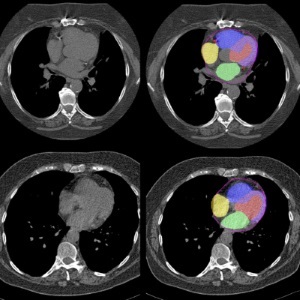
AI Predicts Cardiac Risk and Mortality from Routine Chest CT Scans
Heart disease remains the leading cause of death and is largely preventable, yet many individuals are unaware of their risk until it becomes severe. Early detection through screening can reveal heart issues,... Read moreImaging IT
view channel
New Google Cloud Medical Imaging Suite Makes Imaging Healthcare Data More Accessible
Medical imaging is a critical tool used to diagnose patients, and there are billions of medical images scanned globally each year. Imaging data accounts for about 90% of all healthcare data1 and, until... Read more
Global AI in Medical Diagnostics Market to Be Driven by Demand for Image Recognition in Radiology
The global artificial intelligence (AI) in medical diagnostics market is expanding with early disease detection being one of its key applications and image recognition becoming a compelling consumer proposition... Read moreIndustry News
view channel
Hologic Acquires UK-Based Breast Surgical Guidance Company Endomagnetics Ltd.
Hologic, Inc. (Marlborough, MA, USA) has entered into a definitive agreement to acquire Endomagnetics Ltd. (Cambridge, UK), a privately held developer of breast cancer surgery technologies, for approximately... Read more
Bayer and Google Partner on New AI Product for Radiologists
Medical imaging data comprises around 90% of all healthcare data, and it is a highly complex and rich clinical data modality and serves as a vital tool for diagnosing patients. Each year, billions of medical... Read more